Our work on chloroplast structural haplotype scaffolding has been published in Algorithms for Molecular Biology (BMC)!
From a set of short-read contigs and a set of links between the contigs, we aim to assemble several chloroplast structural haplotypes
https://almob.biomedcentral.com/articles/10.1186/s13015-023-00243-1
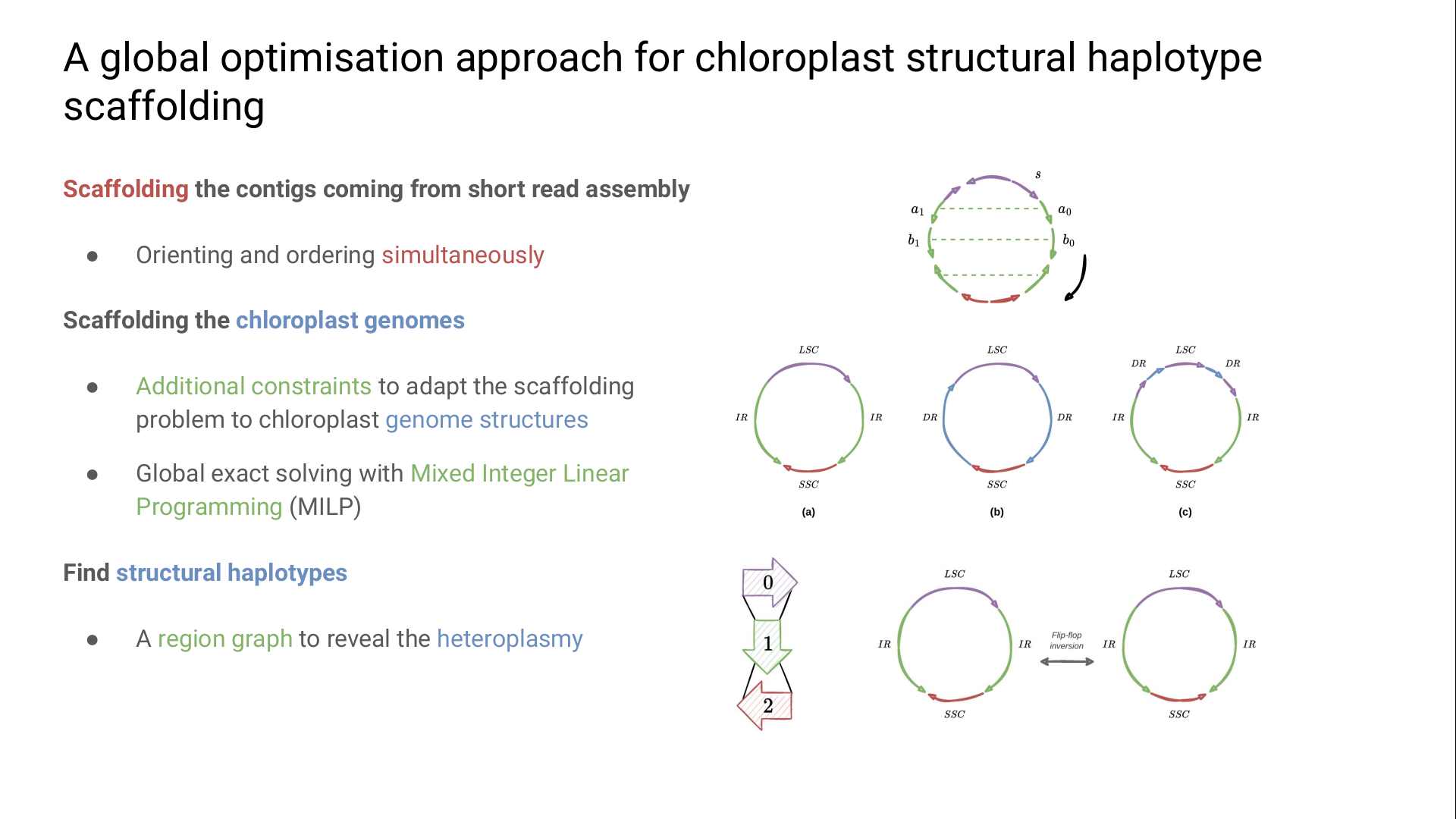
The literature highlights several specific chloroplast circular genomic structures divided into regions, including direct and inverted repeats (DR and IR), and single-copies (SC).
The most studied structure consists of a pair of IR joined with SCs.
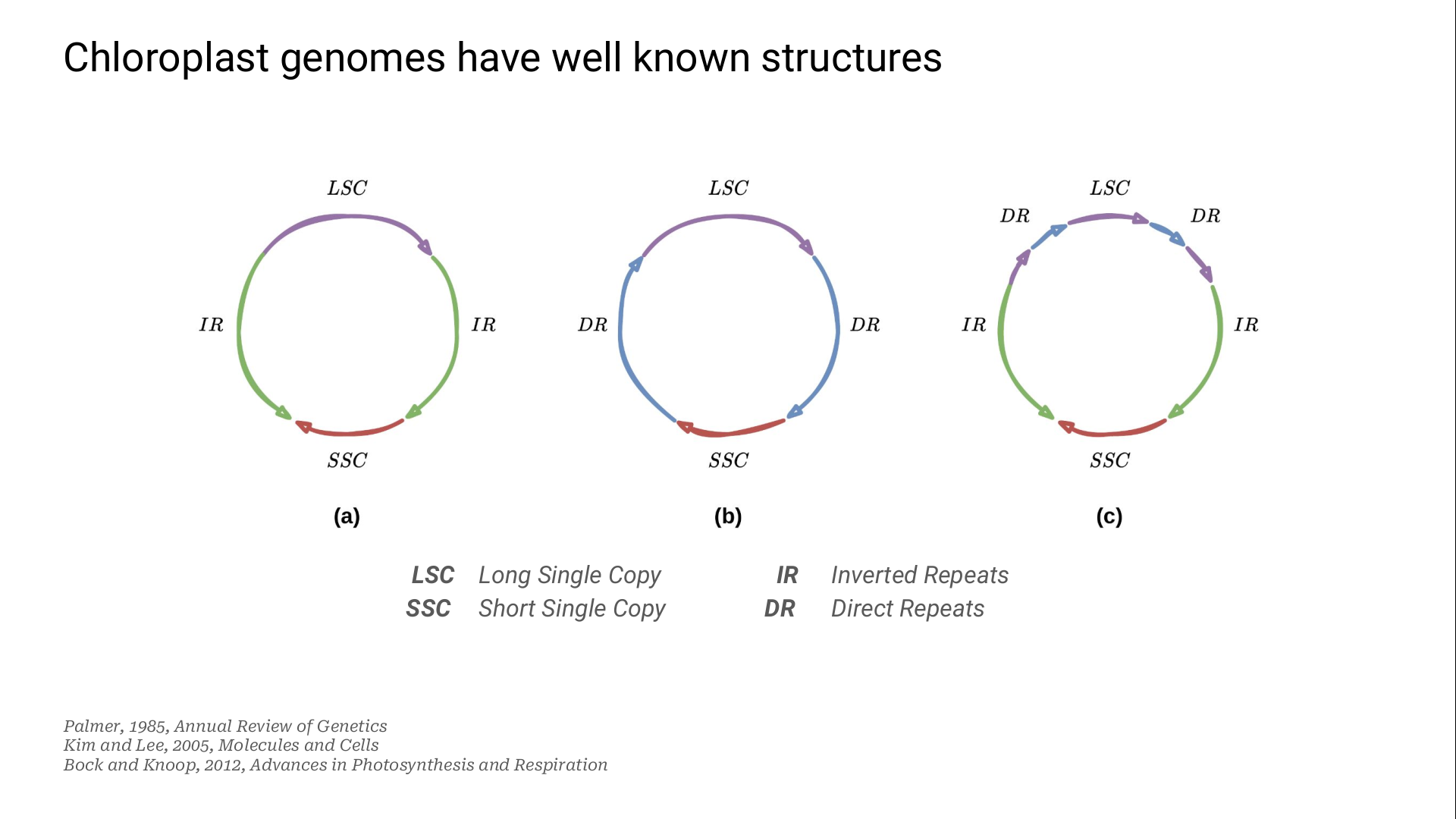
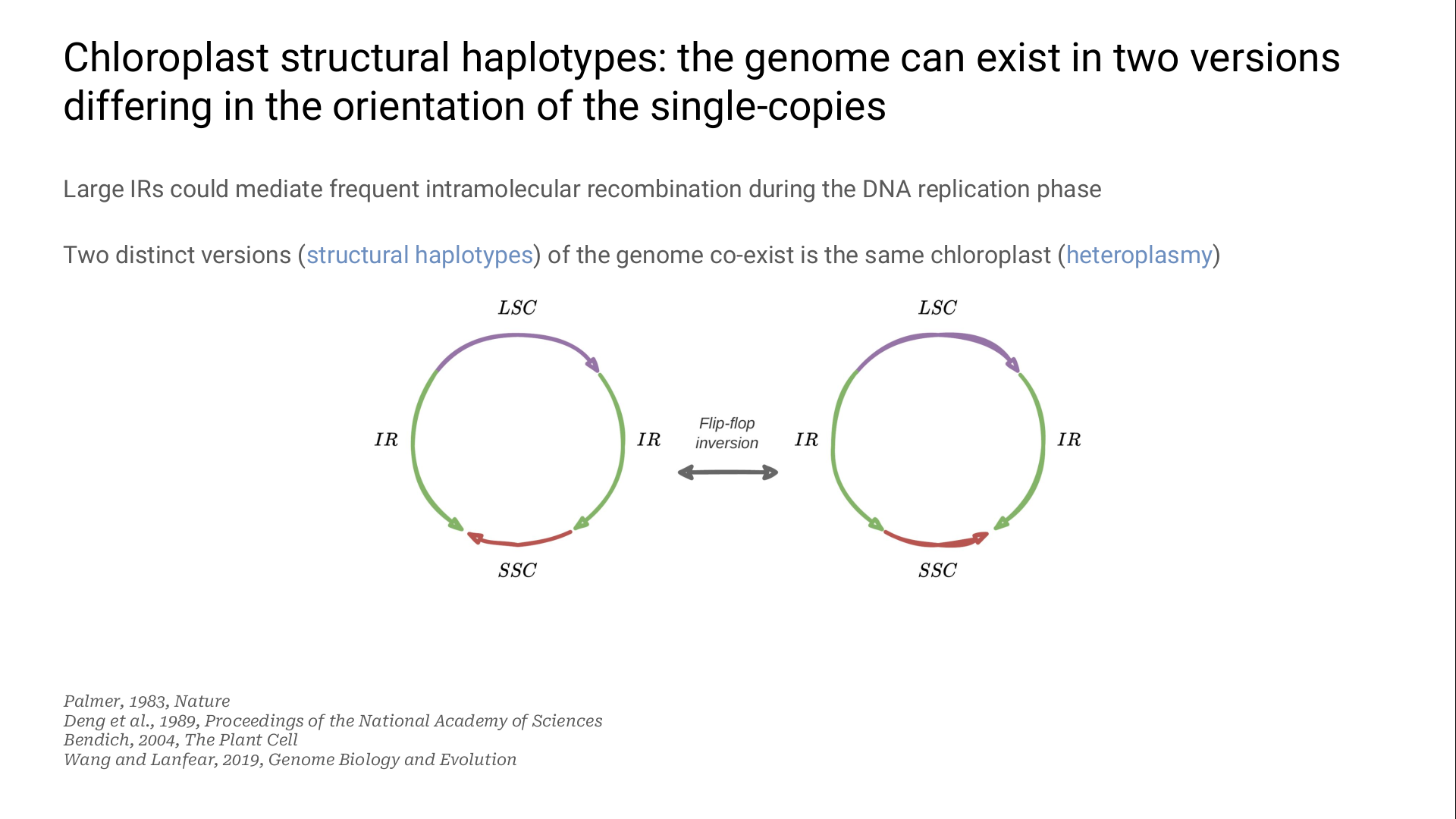
This pair of IR is known to be involved in flip-flop inversions: one of the two SC can be reversed during the DNA replication phase.
These two versions of the genome are structural haplotypes.
They co-exist in the same chloroplast: this phenomenon is named heteroplasmy.
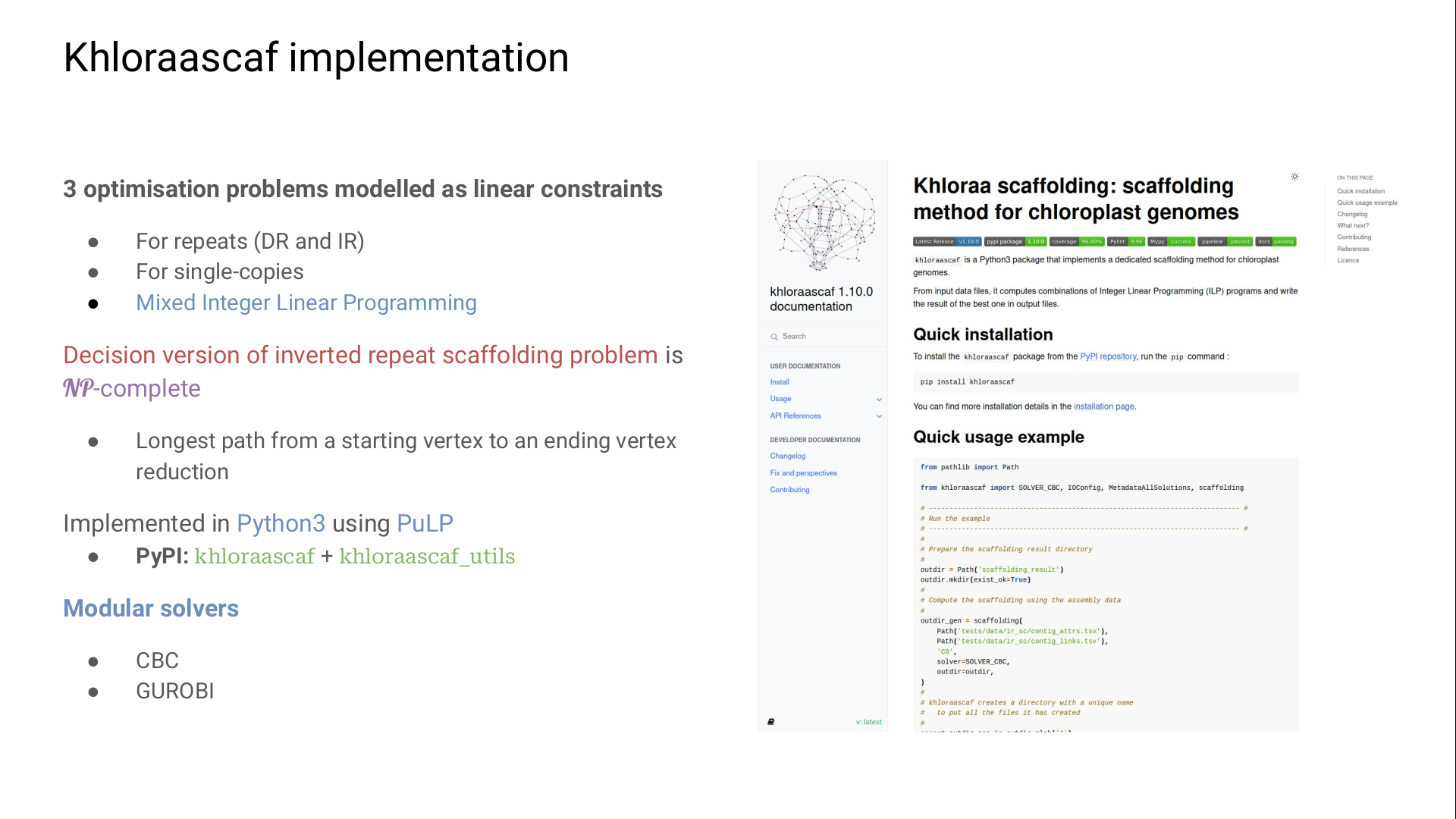
To retrieve several structural haplotypes, our method scaffolds hierarchically each region type.
We first scaffold the repeats and join them by single-copies.
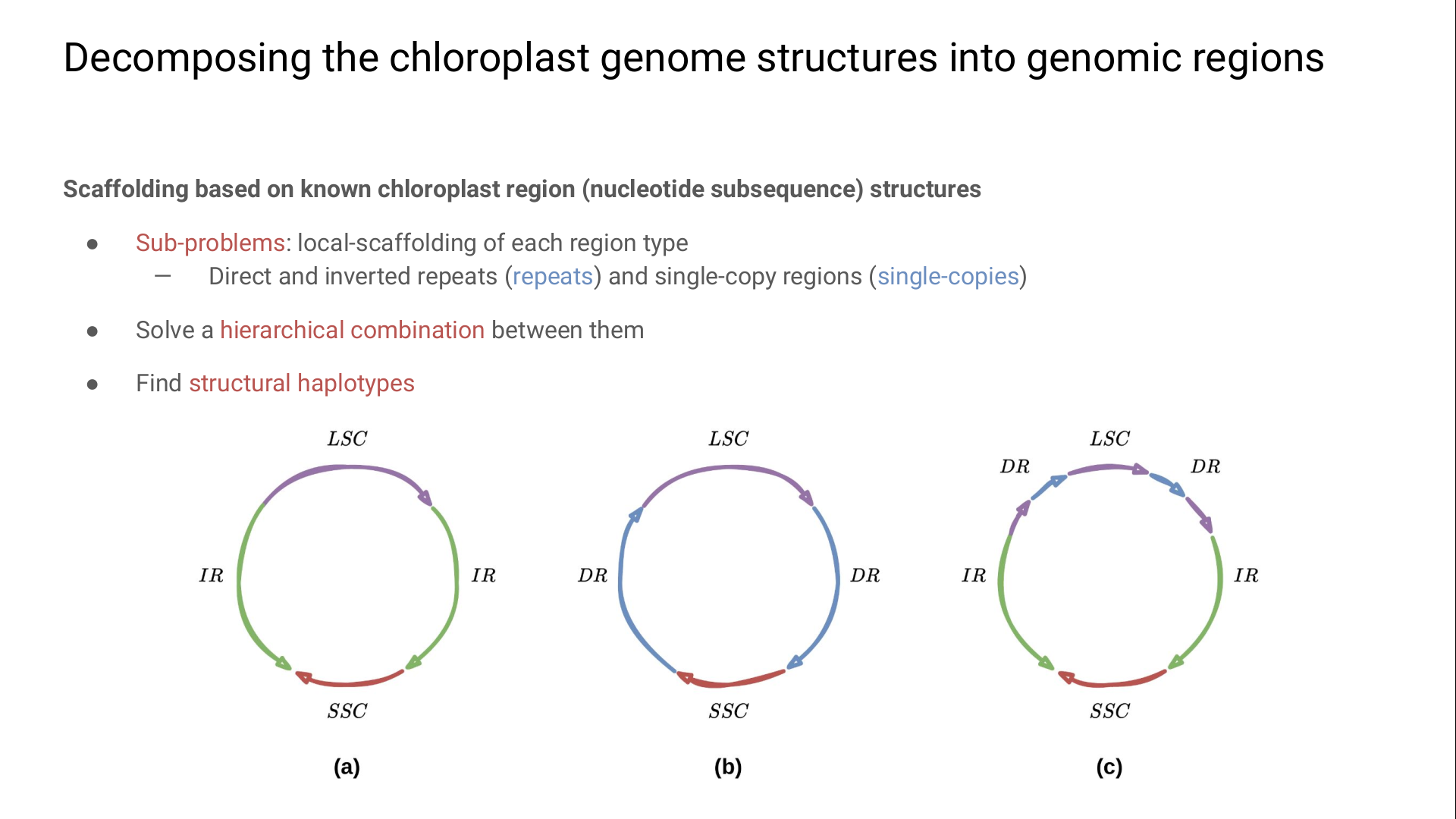

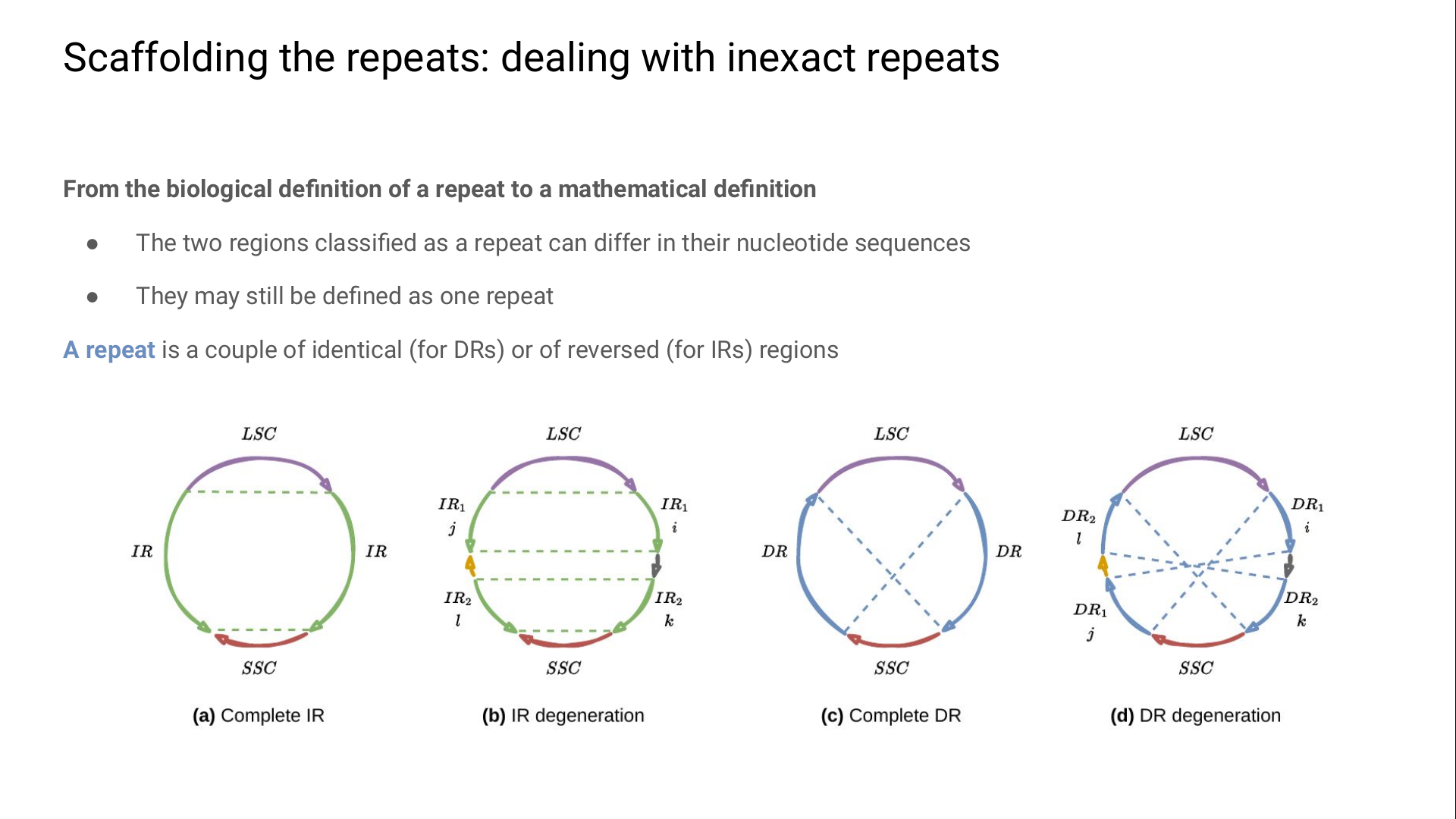
The first tricky thing is to pass from the biological definition of a repeat to one we can exploit mathematically:
here we define a repeat as a couple of identical (for DRs) or perfectly-reversed regions (for IRs).
In our application case, the contigs come from short-read data.
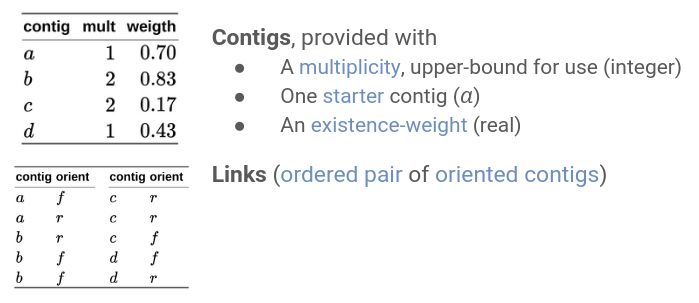
Each contig is provided with a multiplicity (an upper-bound for use) and an existence-weight.
We also have a contig that is known to be part of a single-copy (multiplicity = 1).
The links are ordered pairs of oriented contigs.

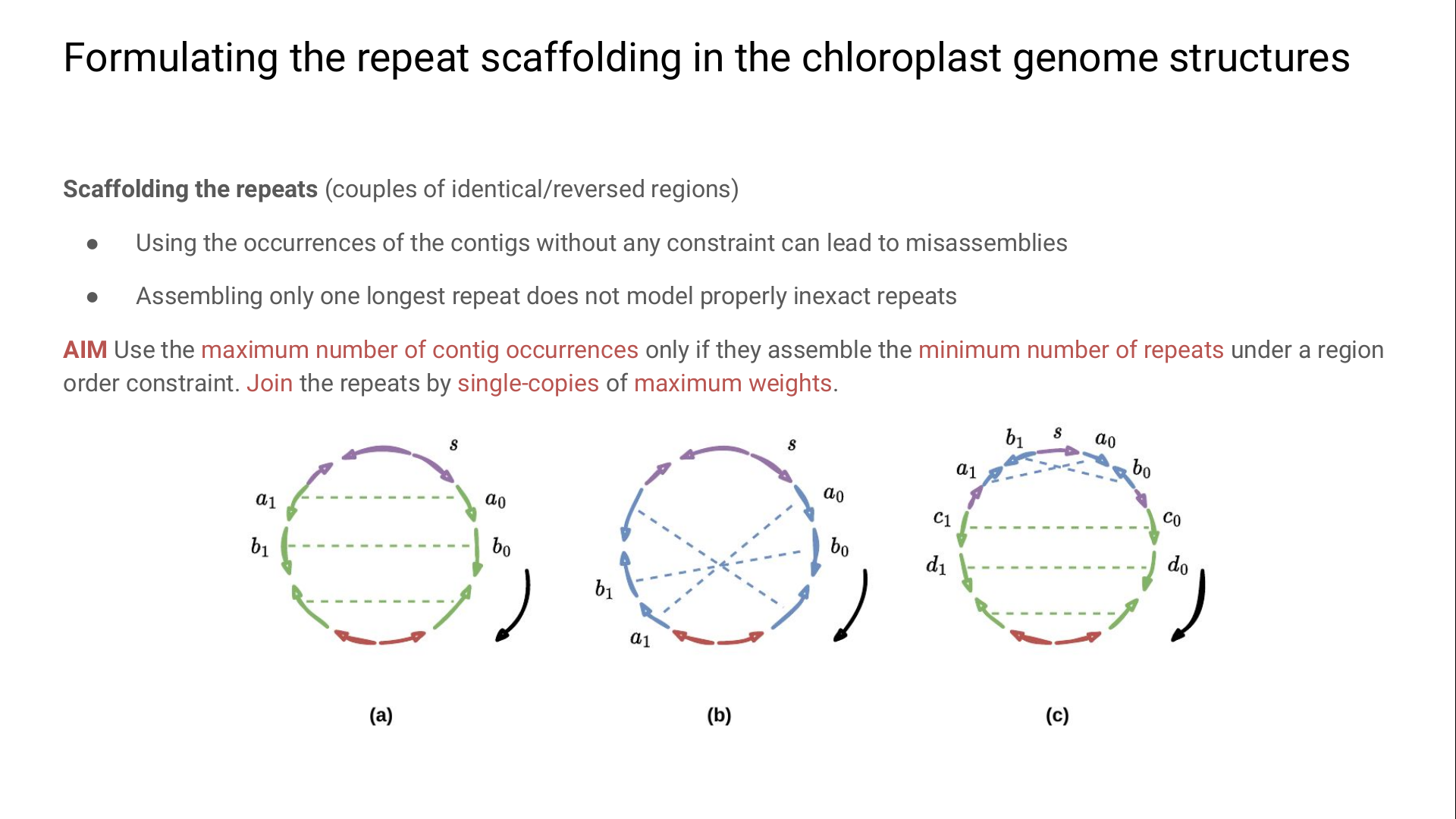
We define the chloroplast scaffolding problem (CHSP) as:
- using the maximum number of contig occurrences only if they assemble the minimum number of repeats under a region order constraint;
- joining the repeats by single-copies of maximum weights.
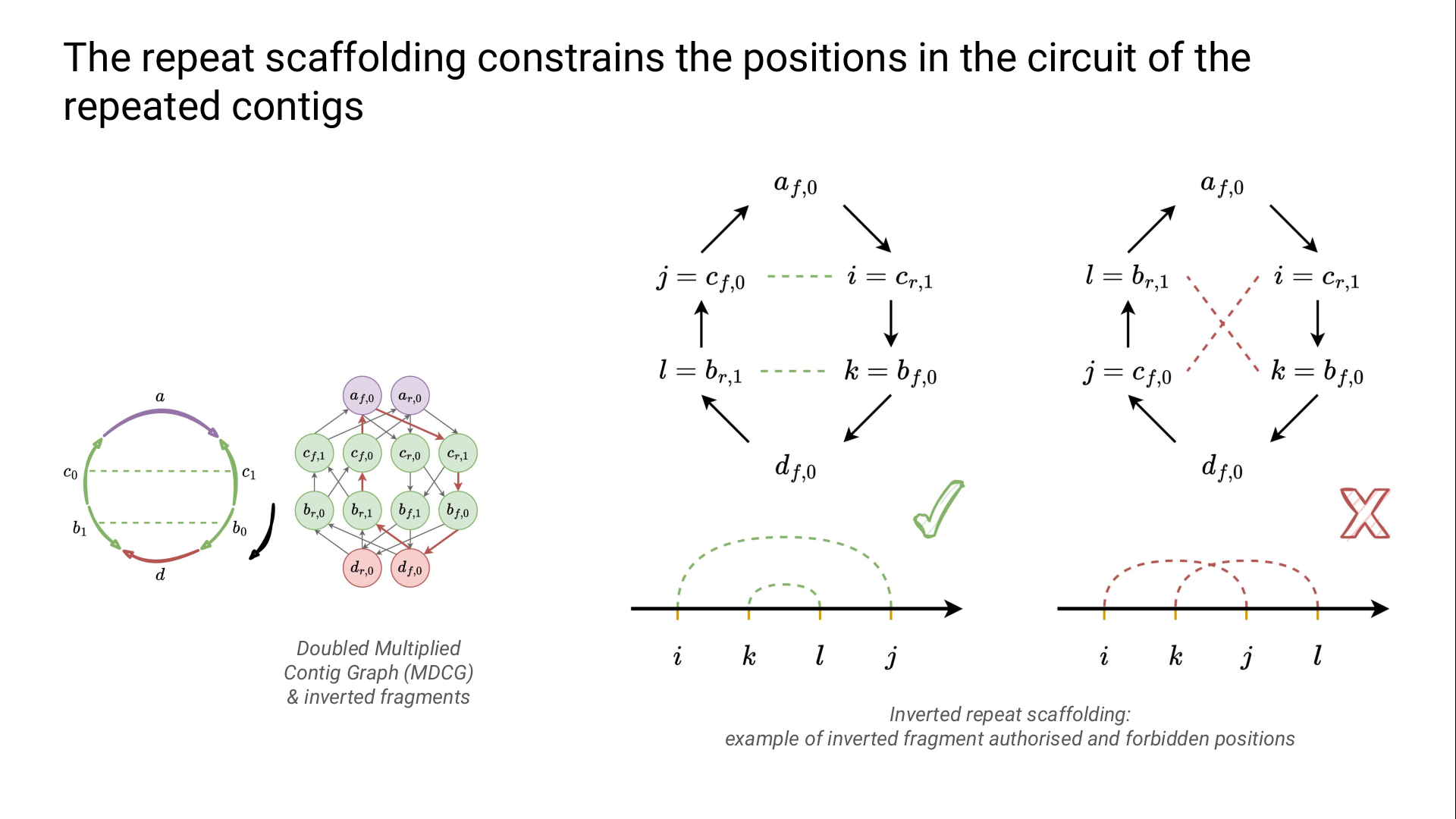
The contigs and their links are represented in a directed fragment graph.
For each contig, there is one vertex for its two possible orientation (forward/reverse).
The genome is a circuit in this graph.
We model the region order constraints for the DRs and the IRs thanks to ILPs.
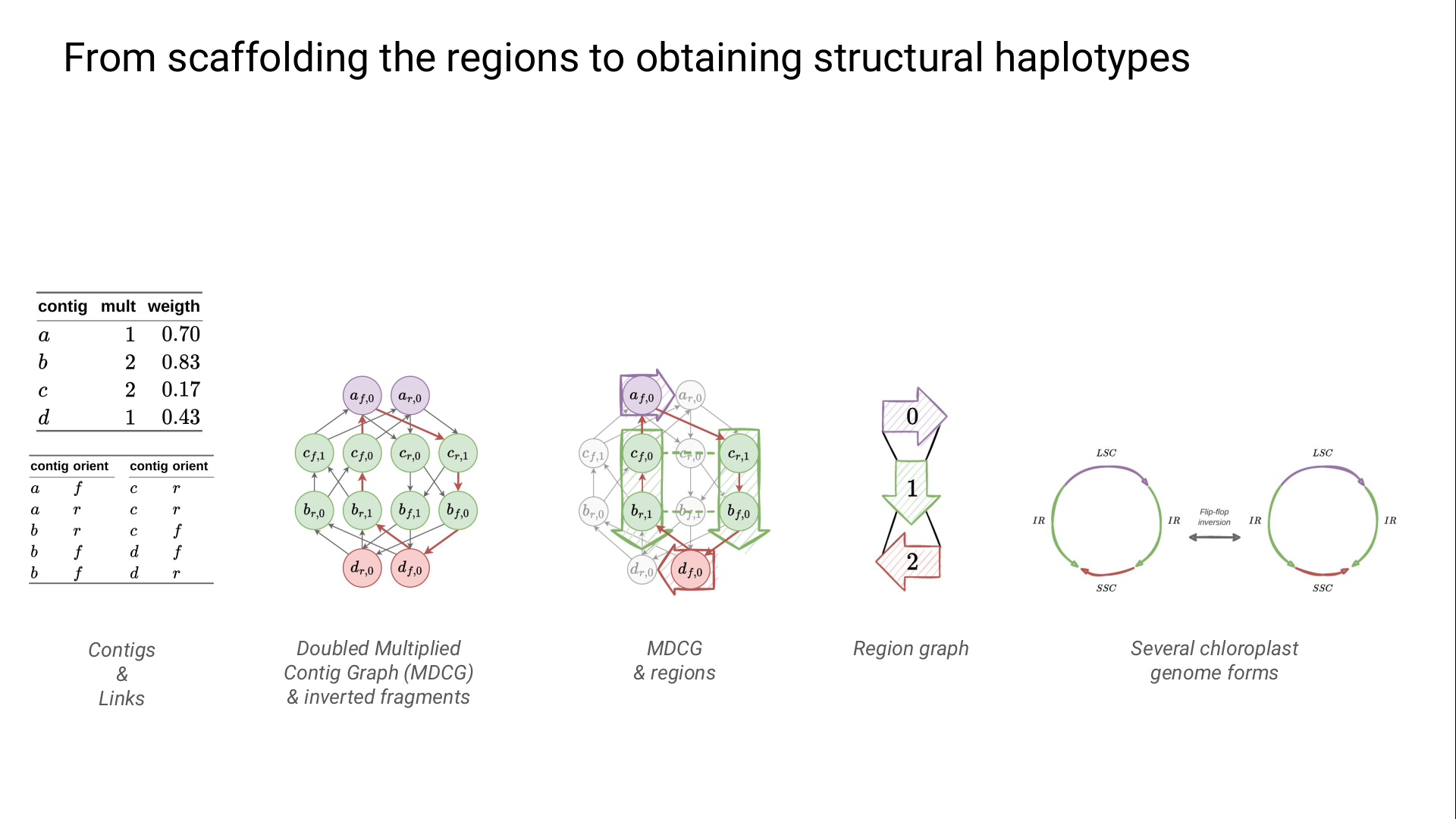
For each region type scaffolding, we fix previously scaffolded regions.
When no region type remains, we extract the regions from the final circuit and represent them in a region graph.
An Eulerian circuit in this last graph corresponds to a structural haplotype.
We have tested our approach with synthetic contigs coming for various chloroplast genome structures and get very encouraging results.
Interestingly, the results suggest to not only one optimal solution but a pool of near-optimal solutions.

The method is available through the khloraascaf Python PyPI package: https://pypi.org/project/khloraascaf/.
The codes to run the tests are available, c.f. https://khloraascaf-results.readthedocs.io/en/latest/.